
When infection rages and other antibiotics prove ineffective, healthcare providers may turn to colistin as their weapon of last resort. But disease-causing microbes are rapidly unraveling the secrets of colistin, and the drug may not be an effective fallback for long.
Researchers at Cornell University have found a ninth variation of a gene that confers resistance to colistin, this time in Salmonella enterica. Called mcr-9, short for mobilized colistin-resistance, this genetic variation (homolog) hardens disease-causing bacteria against the effects of the antibiotic-of-last-resort colistin, just like its cousins mcr-1 through -8.
Here’s why that has researchers scrambling to find a solution.
What Is Colistin?
Colistin combatted Gram-negative bacterial infections such as E. coli, salmonella and shigella from the 1950s through the 80s. It fell out of favor due to its toxic effects on the kidneys, which can lead to chronic disease and renal failure. But it’s now seeing a renaissance, thanks to antibiotic resistance.
Colistin belongs to a group of antibiotics known as polymyxins. Polymyxins like colistin are a class of antibiotics derived from P. polymyxa, a bacterium used as a soil inoculant in agriculture and horticulture. P. polymyxa secretes a substance that protects plants’ roots from disease.
How Colistin Works
Colistin acts like a battering ram or saboteur. Its job is to breach the cell membranes of Gram-negative bacteria. The drug homes in on a certain fatty acid—lipid A— found on the outer cell membrane of certain Gram-negative bacteria and binds to it. This destabilizes the fatty acid, which kicks off a chain reaction:
- The cell membrane expands and cracks in places
- Cellular fluid leaks out
- The pressure inside the cell, or osmotic balance, changes
- The bacterial cell begins to collapse and eventually dies
Colistin is often used in combination therapies with other antibiotics. The breaches that colistin opens in the cell membrane also allow antibiotics such as carbapenems, tetracyclines and others to rush into the cell and destroy it faster.
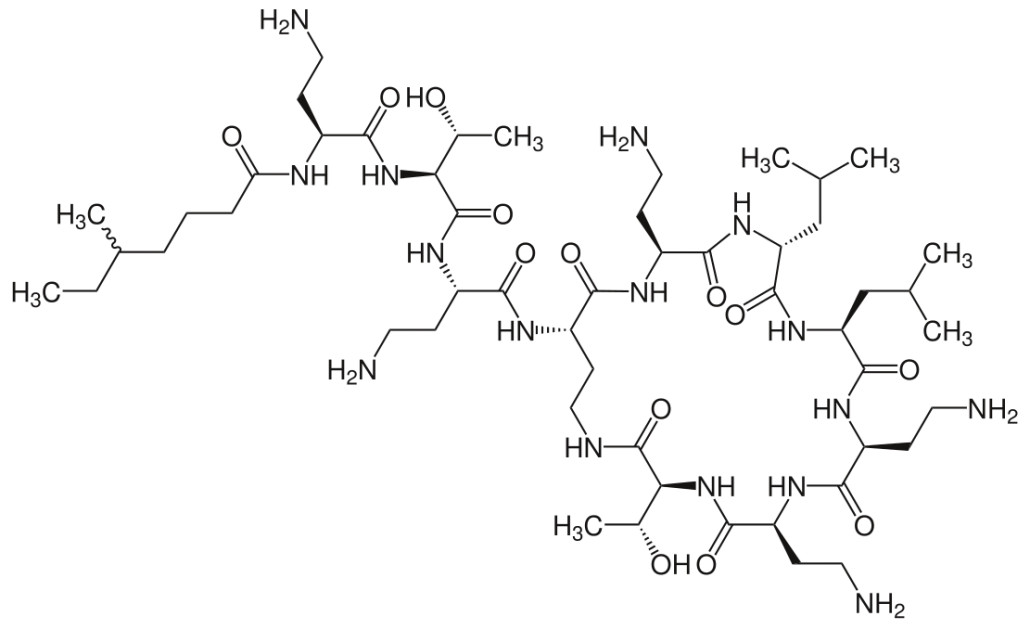
Colistin’s Targets
Healthcare providers deploy colistin to help some of their sickest patients. Researchers have investigated using colistin to fight multidrug-resistant infections in people with cancer, people with cystic fibrosis and people who need a ventilator to breathe for them.
Lung infections are a frequent target of colistin, since the drug can be administered as a nasal spray. Colistin fights both some of the most common disease agents like E. coli and salmonella and some of the most dangerous. It can be effective against multidrug-resistant A.baumannii, P. aeruginosa and enterobacteriaceae like K. pneumoniae, all of which top the World Health Organization’s list of critical threat pathogens.
The MCR Gene
One of healthcare’s most powerful weapons in the war on antibiotic resistance, colistin, might be compromised even now. Some bacteria are developing a secret weapon of their own. Actually, it’s more like armor.
The mcr genes contain instructions for a protein that coats the lipid A on the outer membrane of bacteria cells with a resin that acts like grease. Colistin molecules slide off the lipid A instead of binding to it. Unable to breach the walls, colistin is rendered ineffective.
How Antibiotic Resistance Spreads
For years it was thought that genes that granted resistance to colistin formed only through chromosomal mutations. That is, until mcr-1 was discovered in E. coli in 2015.
MCR genes are formed by plasmids. Plasmids are rogue strands of DNA, not quite alive, that exist in a sort of symbiotic relationship with bacterial cells. Plasmids, like chromosomes, can contain genes and can replicate, often bestowing survivability boosts to their bacterial hosts in the form of genes beneficial to the bacterium, such as antibiotic resistance.
Through a process called horizontal gene transfer, individual bacteria can acquire plasmids and their genes from other bacteria. What’s more, plasmids can transfer between bacteria of different types. MCR genes have been found in:
- E. coli
- Citrobacter
- Enterobacter
- K. pneumoniae
- Salmonella
…and others. Horizontal gene transfer allows antibiotic-resistant genes to jump from bacteria that affect animals to those that affect humans.
Many researchers dread the thought of finding mcr genes in the so-called nightmare bacteria, carbapenem-resistant Enterobacteriaceae (CRE). These bugs are already resistant to carbapenem, itself an antibiotic-of-last-resort. Currently, colistin is one of the only effective treatments for CRE infections.
Global Action
If there’s one bright spot to the emergence of mcr genes, it’s that in less than five years from the discovery of mcr-1, eight homologs have been identified. It shows that the scientific and medical community as a whole is committed to battling antibiotic resistance on all fronts.
Antibiotic resistance is broadly recognized as one of the greatest public health threats the world has seen. Responsible use of antibiotics is crucial to slowing its spread, ensuring common infections don’t turn deadly and preventing a post-antibiotic world.
Policy and Monitoring
India’s Health Ministry in July banned the use of colistin in farm animals. Overreliance on antibiotics for livestock is one of the top reasons for the spread of antibiotic resistance. And, thanks to plasmids, bacteria in animals can transfer drug resistance to bacteria in humans, so it’s especially important to use antibiotics responsibly.

In July 2019, mcr-3 was found in a bacterial sample taken in 2010 from a human, an 18-year-old North Carolina resident. Only through vigilant monitoring programs, such as that of the Centers of Disease Resistance and Prevention, can we take the first steps in avoiding a colistin resistance ambush.
Luckily, dedicated researchers across the world are rushing to identify mcr genes, and hopefully, find a workaround. In the meantime, the war between humans and bacterial infection rages on.

